 Batched Imputation for High Dimensional Missing Data Problems.How to use Lowess Smoothing in R with example.This is the algorithm of Holm step-down procedure. If it is not make sure it is equal to the largest of the preceding p-values. We then insure that any adjusted p-value is at least as large as any preceding adjusted p-value. Compare each of these adjusted p-values to 0.05. Multiply the third smallest p-value by 998, the fourth smallest by 997, etc. Then for the 2nd one, multiply its p-value by 999 (not one thousand) and see if it is less than 0.05. There is no difference as Bonferroni test for the gene. If that adjusted p-value is less than 0.05, then that gene shows evidence of differential expression. Multiply the smallest p-value by one thousand. First, sort your thousand p-values from low to high. The Holm step-down procedure is the easiest to understand. P.adjust(p, method = "bonferroni")The sequential corrections is slightly more powerful than Bonferroni test. So, when doing corrections, simply multiply the nominal p-value by m to get the adjusted p-values. It rejects any hypothesis with p-value ≤ α/m. Bonferroni correction is to control the overall type I errors when all tests are independent. Bonferroni) and sequential adjustment (e.g. Holm or Hochberg). It consists of two types : single-step (e.g. Many procedures have been developed to control the family-wise error rate P(V≥ 1), including the Bonferroni, Holm (1979), Hochberg (1988), and Sidak. Positive false discovery rate (pFDR): the rate that discoveries are false, pFDR = E(V/R | R > 0)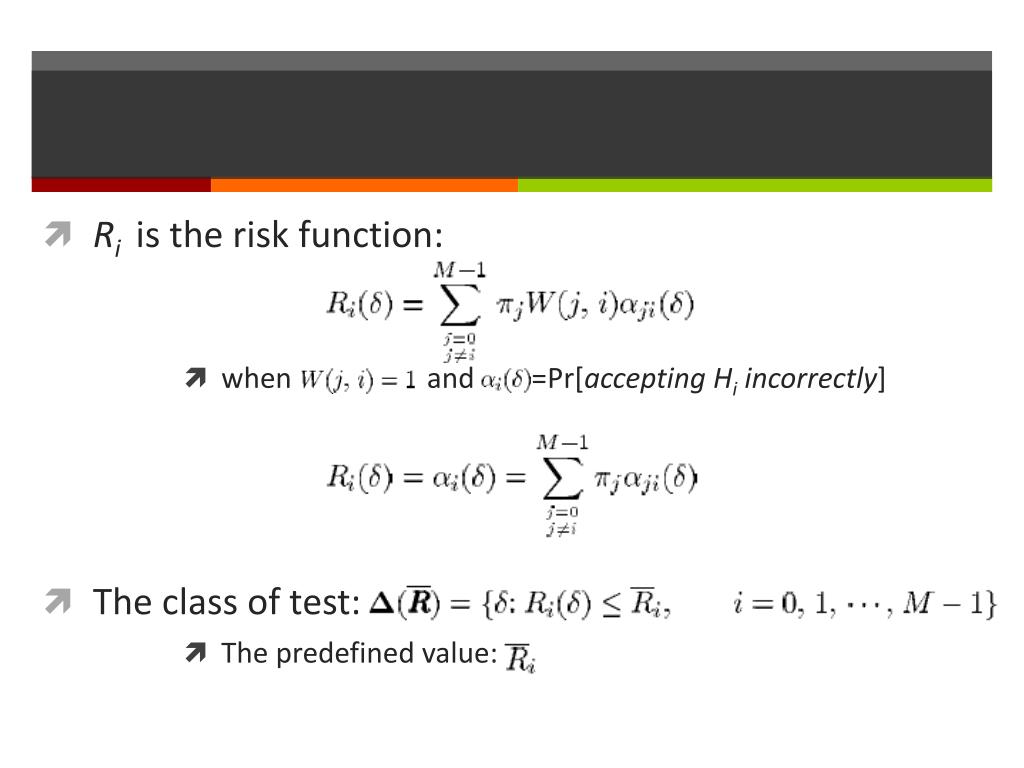 
Per-family error rate (PFER): the expected number of Type I errors, PFE = E(V).įamily-wise error rate (FWER): the probability of at least one type I error, FWER = P(V ≥ 1)įalse discovery rate (FDR) is the expected proportion of Type I errors among the rejected hypotheses, FDR = E(V/R | R>0)P(R>0) Per comparison error rate (PCER): the expected value of the number of Type I errors over the number of hypotheses, PCER = E(V)/m There are many different ways to control the type I errors, such as If your chance of making an error in single test is α, then your chance to make one or more errors in m tests will be It matters because we usually perform the same hypothesis tests not just once, but many many times. – Sensitivity and Specificity Total test (m) Power of a test = the ability to detect True Positive among all real positive cases. So when a single test reaches p-value 0.05, we can intuitively understand that with 5% of chance we make a mistake or 5% of cases we thought significant are actually not. β is related with the power of a test. The similar logics for Type II error, or false negative.Īlso note that people use Greek letter α for type I error rate and β for type II error rate. α is also the significant level for a test, e.g. no difference, no effect), then you reject it (or yes, there is difference), it’s like a false positive. If the true thing is a null hypothesis (H0), which is what people usually assume (e.g. 
Type I error is “you reject a true thing”. 
When we do a hypothesis test, we can categorize the result into the following 2×2 table: – Type I error (false positive) and Type II error (false negative): Multiple testing is such a piece in my knowledge map. If we like to bound these over-fitting probabilities by α, then we can assign different values for each α(k a,k b).It’s not a shame to put a note on something (probably) everyone knows and you thought you know but actually you are not 100% sure. 
If the null is rejected, then increase k a by 1 otherwise, decrease k b by 1. The procedure begins with k a= K min and k b= K max. Each one tests the null hypothesis H 0: k = k a against the alternative hypothesis H 1: k = k b. The Joinpoint program uses a sequence of "permutation" tests to select the final model. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
